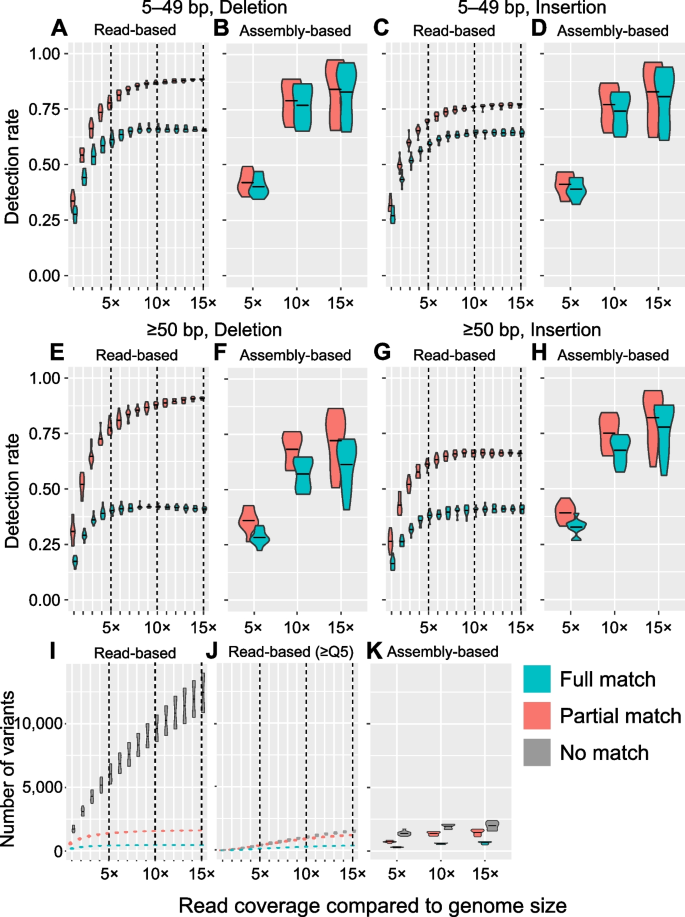
Benchmarking datasets for assembly-based variant calling using high-fidelity long reads | BMC Genomics | Full Text
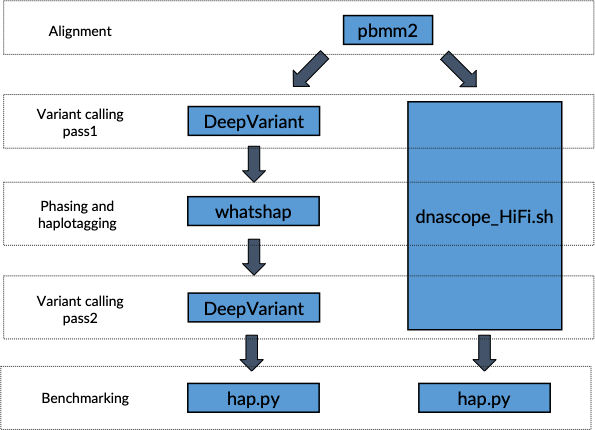
An exploration of machine-learning based variant callers for SNV and small-indel using PacBio HiFi data | by Yih-Chii Hwang, Ph.D. | DNAnexus Science Frontiers | Medium






















